InvariantCausal.jl
High-performance implementation of invariant causal prediction
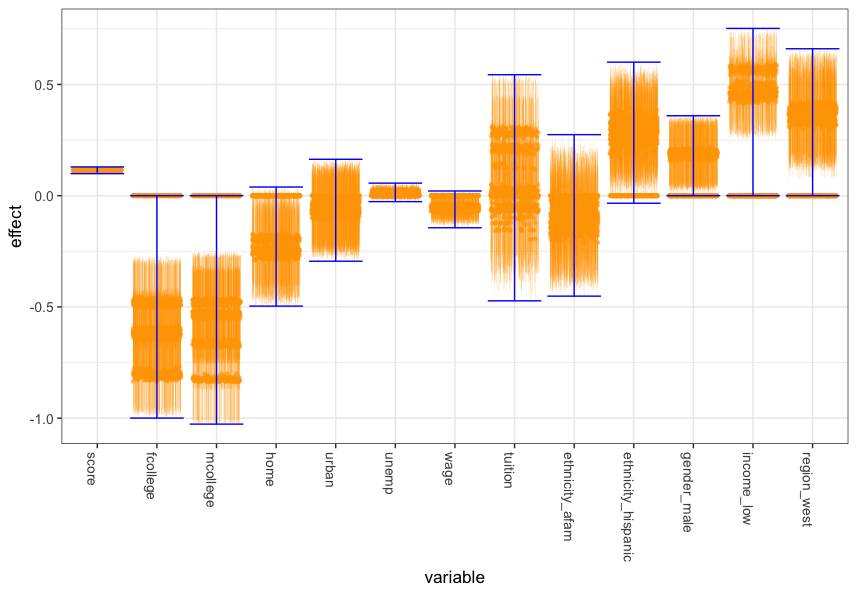
This is a Julia 1.x implementation for the Invariant Causal Prediction algorithm of Peters, Bühlmann and Meinshausen. The method uncovers direct causes of a target variable from datasets under different environments (e.g., interventions or experimental settings).